Role of Post-Translational Modifications in Pathogenesis of Rheumatoid Arthritis and Therapeutic Targeting: A Comprehensive Systematic Review
Keywords:
Rheumatoid Arthritis, Protein Processing, Post-Translational Modifications, Phosphorylation, Ubiquitination, Acetylation, Autoimmunity, Signal Transduction, CytokineAbstract
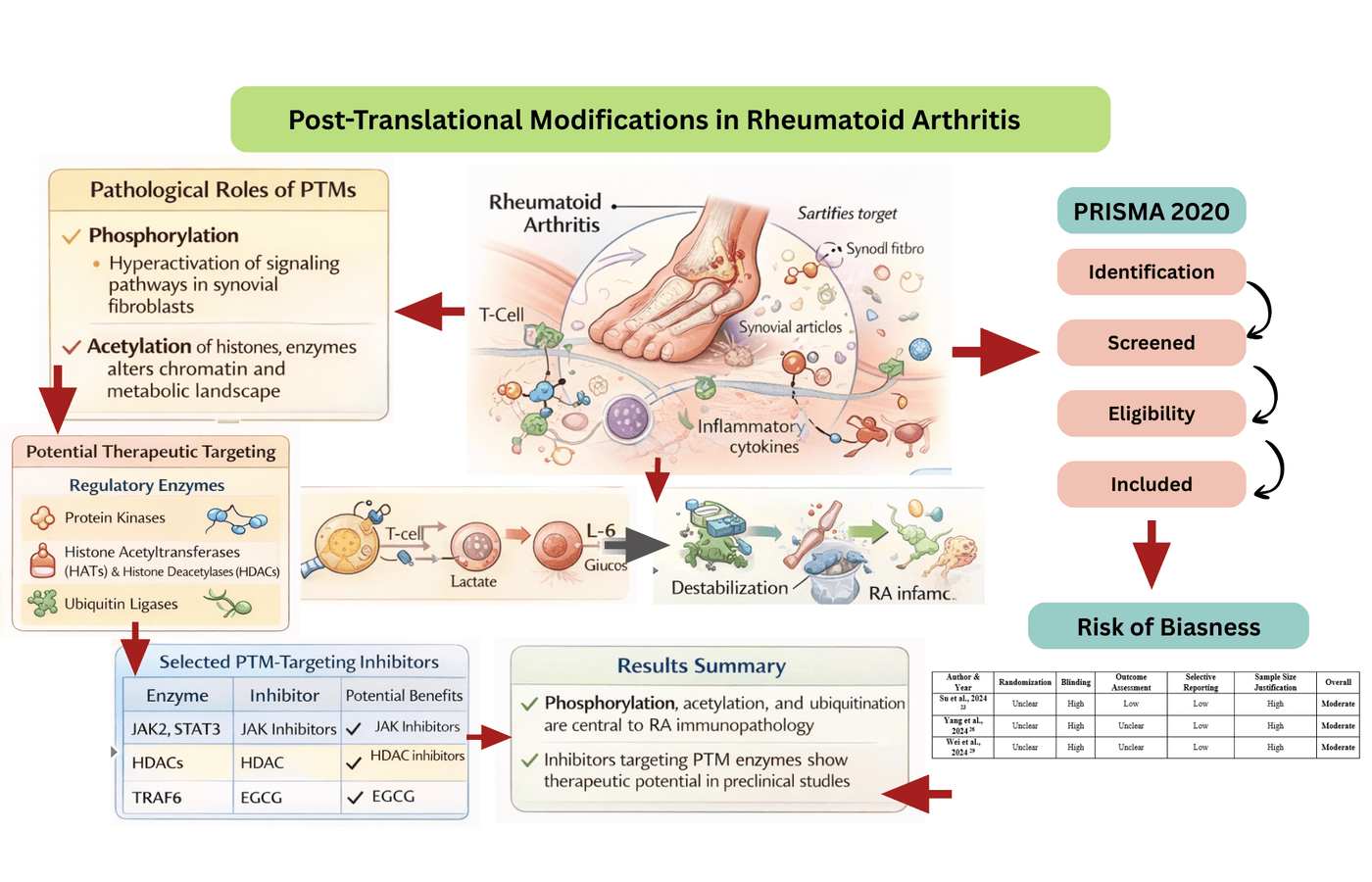
Background: Post-translational modifications (PTMs) such as phosphorylation, acetylation, and ubiquitination played a critical role in pathogenesis of inflammatory diseases including rheumatoid arthritis (RA). This systematic review aimed to synthesize evidence of principal role of PTMs in RA and to evaluate the therapeutic potential of agents targeting these dysregulated modification pathways. Methods: This systematic review followed PRISMA guidelines 2020. Research was conducted in PubMed, Scopus, Web of Science, and Google Scholar from 2015 to 2025. The studies which explored the role of particular PTMs in the RA pathophysiology and their therapeutic potential, were considered. Articles and reviews that were not written in English and where mechanistic or therapeutic outcomes were not described were excluded. The JBI Critical Appraisal Checklist, Newcastle-Ottawa Scale, and SYRCLE tool were used for evaluating risk of bias whereas, quality of study was evaluated using GRADE approach. Results: The findings of twelve studies demonstrated that dysregulation of PTMs played a substantial role in RA immunopathology. Mechanistically, acetylation, ubiquitination, and phosphorylation are involved in dysregulated T-cell metabolism, Th17/ Treg imbalance, synovial fibroblast activation, and pro-inflammatory signaling. Preclinical studies supported the therapeutic potential of targeting associated regulatory enzymes. The risk of bias of most studies was measured as moderate and certainty of evidence was found to be low. Conclusion: Phosphorylation, ubiquitination, and acetylation played a critical role in the pathogenesis of RA and could be exploited as potential therapeutic targets.
References
1. Jahid M, Khan KU, Ahmed RS. Overview of rheumatoid arthritis and scientific understanding of the disease. Mediterr J Rheumatol. 2023;34(3):284-91. https://doi.org/10.31138/mjr.20230801.oo
2. Alivernini S, Firestein GS, McInnes IB. The pathogenesis of rheumatoid arthritis. Immunity. 2022;55(12):2255-70. https://doi.org/10.1016/j.immuni.2022.11.009
3. Babaahmadi M, Tayebi B, Gholipour NM, Kamardi MT, Heidari S, Baharvand H, et al. Rheumatoid arthritis: the old issue, the new therapeutic approach. Stem cell res ther. 2023;14(1):268. https://doi.org/10.1186/s13287-023-03473-7
4. Mnatsakanyan R, Shema G, Basik M, Batist G, Borchers CH, Sickmann A, et al. Detecting post-translational modification signatures as potential biomarkers in clinical mass spectrometry. Expert Rev Proteomics. 2018;15(6):515-35. https://doi.org/10.1080/14789450.2018.1483340
5. Mastrangelo A, Colasanti T, Barbati C, Pecani A, Sabatinelli D, Pendolino M, et al. The role of posttranslational protein modifications in rheumatological diseases: focus on rheumatoid arthritis. J Immunol Res. 2015;2015(1):712490. http://dx.doi.org/10.1155/2015/712490
6. Willems P, Sterck L, Dard A, Huang J, De Smet I, Gevaert K, et al. The Plant PTM Viewer 2.0: in-depth exploration of plant protein modification landscapes. J Exp Bot. 2024;75(15):4611-24. https://doi.org/10.1093/jxb/erae270
7. Wen J, Wang D. Deciphering the PTM codes of the tumor suppressor p53. J Mol Cell Biol. 2021;13(11):774-85. https://doi.org/10.1093/jmcb/mjab047
8. Stastna M. Post‐translational modifications of proteins in cardiovascular diseases examined by proteomic approaches. The FEBS J. 2025;292(1):28-46. https://doi.org/10.1111/febs.17108
9. Zhong Q, Xiao X, Qiu Y, Xu Z, Chen C, Chong B, et al. Protein posttranslational modifications in health and diseases: Functions, regulatory mechanisms, and therapeutic implications. MedComm. 2023;4(3):e261. https://doi.org/10.1002/mco2.261
10. Safa A, Vruzhaj I, Gambirasi M, Toffoli G. Antigenic dark matter: unexplored post-translational modifications of tumor-associated and tumor-specific antigens in pancreatic cancer. Cancers. 2025;17(21):3506. https://doi.org/10.3390/cancers17213506
11. Haro I, Sanmartí R, Gómara MJ. Implications of post-translational modifications in autoimmunity with emphasis on citrullination, homocitrullination and acetylation for the pathogenesis, diagnosis and prognosis of rheumatoid arthritis. Int J Mol Sci. 2022;23(24):15803. https://doi.org/10.3390/ijms232415803
12. Sun C-F, Li Y-Q, Mao X-M. Regulation of protein post-translational modifications on metabolism of actinomycetes. Biomolecules. 2020;10(8):1122. https://doi.org/10.3390/biom10081122
13. Ortiz AM, Silva Fernández L, Villaverde V, Abad MÁ, Maneiro JR, Candelas G, et al. Gaps between research and recommendations in rheumatoid arthritis. Rheumatol Int. 2021;41(1):57-66. https://doi.org/10.1007/s00296-020-04724-z
14. Page MJ, McKenzie JE, Bossuyt PM, Boutron I, Hoffmann TC, Mulrow CD, et al. The PRISMA 2020 statement: an updated guideline for reporting systematic reviews. bmj. 2021;372. https://doi.org/10.1136/bmj.n71
15. Barker TH, Stone JC, Sears K, Klugar M, Tufanaru C, Leonardi-Bee J, et al. The revised JBI critical appraisal tool for the assessment of risk of bias for randomized controlled trials. JBI evidence synthesis. 2023;21(3):494-506. https://doi.org/10.11124/JBIES-22-00430
16. Hooijmans CR, Rovers MM, De Vries RB, Leenaars M, Ritskes-Hoitinga M, Langendam MW. SYRCLE’s risk of bias tool for animal studies. BMC Med Res Methodol. 2014;14(1):43. https://doi.org/10.1186/1471-2288-14-43
17. Carra MC, Romandini P, Romandini M. Risk of bias evaluation of cross-sectional studies: Adaptation of the Newcastle–Ottawa Scale. J Periodontal Res. 2025 Apr;60(2):e13405. https://doi.org/10.1111/jre.13405
18. Haro I, Castellanos-Moreira R, Sanmartí R, Gómara MJ. Anti-Modified Peptide Antibodies (AMPAs) in Rheumatoid Arthritis: Study of the Diagnostic Value of Citrullinated, Homocitrullinated, and Acetylated Fibrin/Filaggrin Chimeric Peptides. Diagnostics. 2024;14(22):2485. https://doi.org/10.3390/diagnostics14222485
19. Arito M, Nagai K, Ooka S, Sato T, Takakuwa Y, Kurokawa M, et al. Altered acetylation of proteins in patients with rheumatoid arthritis revealed by acetyl-proteomics. Clin Exp Rheumatol. 2015;33(6):877-86. https://doi.org/10.55563/clinexprheumatol/8d0gx3
20. Nijjar JS, Morton FR, Bang H, Buckley CD, van der Heijde D, Gilmour A, et al. The impact of autoantibodies against citrullinated, carbamylated, and acetylated peptides on radiographic progression in patients with new-onset rheumatoid arthritis: an observational cohort study. Lancet Rheumatol. 2021;3(4):e284-e93. https://doi.org/10.1016/S2665-9913(20)30381-7
21. García-Moreno C, Gómara MJ, Castellanos-Moreira R, Sanmartí R, Haro I. Peptides bearing multiple post-translational modifications as antigenic targets for severe rheumatoid arthritis patients. Int J Mol Sci. 2021;22(24):13290. https://doi.org/10.3390/ijms222413290
22. Wang W, Xiang T, Yang Y, Wang Z, Xie J. E3 ubiquitin ligases STUB1/CHIP contributes to the Th17/Treg imbalance via the ubiquitination of aryl hydrocarbon receptor in rheumatoid arthritis. Clin Exp Immunol. 2022;209(3):280-90. https://doi.org/10.1093/cei/uxac072
23. Su X, Wang X, Zhang X, Sun Y, Jia Y. β-Indole-3-acetic acid attenuated collagen-induced arthritis through reducing the ubiquitination of Foxp3 via the AhR-TAZ-Tip60 pathway. Immunologic Research. 2024;72(4):741-53. https://doi.org/10.1007/s12026-024-09480-x
24. Kong Q-Z, Guo L-T, Yang J-N, Wang Y-F, Zhao J-X, Kong S-H, et al. Anti‐inflammatory effects of TRAF‐interacting protein in rheumatoid arthritis fibroblast‐like synoviocytes. Mediators Inflamm. 2016;2016(1):3906108. https://doi.org/10.1155/2016/3906108
25. Singh AK, Umar S, Riegsecker S, Chourasia M, Ahmed S. Regulation of transforming growth factor β–activated kinase activation by epigallocatechin‐3‐gallate in rheumatoid arthritis synovial fibroblasts: suppression of K63‐linked autoubiquitination of tumor necrosis factor receptor–associated factor 6. Arthritis & Rheumatology. 2016;68(2):347-58. https://doi.org/10.1002/art.39447
26. Yang G, Liu K, Wang Y, Pan X, Ye J, Li Y, et al. Phosphorylation of NIR‐II emitting Au nanoclusters for targeted bone imaging and improved rheumatoid arthritis therapy. Aggregate. 2024;5(2):e435. https://doi.org/10.1002/agt2.435
27. Zhang X, Zhang D, Wang Q, Guo X, Chen J, Jiang J, et al. Sprouty2 inhibits migration and invasion of fibroblast-like synoviocytes in rheumatoid arthritis by down-regulating ATF2 expression and phosphorylation. Inflammation. 2021;44(1):91-103. https://doi.org/10.1007/s10753-020-01311-z
28. Novak-Kotzer H, Capera J, Jainarayanan A, Elanchezhian M, Valvo S, Mayya V, et al. STAT3 phosphorylation in the rheumatoid arthritis immunological synapse. J Autoimmun. 2025;15(5):103456. https://doi.org/10.1101/2025.01.20.633875
29. Wei Y, Wang D, Wu J, Zhang J. JAK2 inhibitors improve RA combined with pulmonary fibrosis in rats by downregulating SMAD3 phosphorylation. Int J Rheum Dis. 2024;27(5):e15164. https://doi.org/10.1111/1756-185X.15164
30. Zhang G, Yao L, Zhu Q, Herrmann M, Zhao Y. Cross-reactivity of anti-modified protein antibodies in rheumatoid arthritis. Clin Exp Rheumatol. 2024;43(7):1332-44. https://doi.org/10.55563/clinexprheumatol/c0eeg7
31. Trouw LA, Rispens T, Toes RE. Beyond citrullination: other post-translational protein modifications in rheumatoid arthritis. Nat Rev Rheumatol. 2017;13(6):331-9. https://doi.org/10.1038/nrrheum.2017.15
32. Sebastiani M, Manfredi A, Croci S, Faverio P, Cassone G, Vacchi C, et al. Rheumatoid arthritis extra-articular lung disease: new insights on pathogenesis and experimental drugs. Expert Opin Investig Drugs. 2024;33(8):815-27. https://doi.org/10.1080/13543784.2024.2376567
33. Bano U, Memon S, Shahani MY, Shaikh P, Gul S. Epigenetic effects of in utero bisphenol A administration: Diabetogenic and atherogenic changes in mice offspring. Iran J Basic Med Sci. 2019 May;22(5):521-528. https://doi.org/10.22038/ijbms.2019.29909.7357
34. Memon S, Bano U, Shahani MY. Maternal exposure to Bisphenol‐A induces epigenetic changes in metabolic syndrome markers of offspring. FASEB J. 2018 Apr;32(S1):774-2. https://doi.org/10.1096/fasebj.2018.32.1_supplement.774.2
35. Memon S, Bano U, Singha SP, Shahani MY, Shaikh P, Shahani SB, et al. Chromium restriction in the intrauterine environment and development of metabolic diseases in mice offspring. J Taibah Univ Med Sci. 2025;20(3):365–75. https://doi.org/10.1016/j.jtumed.2025.06.001
36. Volkov M, van Schie KA, van der Woude D. Autoantibodies and B Cells: The ABC of rheumatoid arthritis pathophysiology. Immunological reviews. 2020;294(1):148-63. https://doi.org/10.1111/imr.12829
37. Hsieh H-C, Ling L-L, Wang Y-C. Post-translational modifications of immune checkpoints: molecular mechanisms, tumor microenvironment remodeling, and therapeutic implications. J Biomed Sci. 2026;33(1):3. https://doi.org/10.1186/s12929-025-01202-1
38. Li X, Yu T, Li X, He X, Zhang B, Yang Y. Role of novel protein acylation modifications in immunity and its related diseases. Immunology. 2024;173(1):53-75. https://doi.org/10.1111/imm.13822
39. Sasaki K, Iwai K. Role of linear ubiquitination in inflammatory responses and tissue homeostasis. Int Immuno. 2023;35(1):19-25. https://doi.org/10.1093/intimm/dxac047
40. Jiang W, Li M, Peng S, Hu T, Long Y, Zhang J, et al. Ubiquitin ligase enzymes and de-ubiquitinating enzymes regulate innate immunity in the TLR, NLR, RLR, and cGAS-STING pathways. Immunol Res. 2023;71(6):800-13. https://doi.org/10.1007/s12026-023-09400-5
41. Liu J, Cheng Y, Zheng M, Yuan B, Wang Z, Li X, et al. Targeting the ubiquitination/deubiquitination process to regulate immune checkpoint pathways. Signal Transduct Target Ther. 2021;6(1):28. https://doi.org/10.1038/s41392-020-00418-x
42. Maglaviceanu A, Wu B, Kapoor M. Fibroblast‐like synoviocytes: role in synovial fibrosis associated with osteoarthritis. Wound Repair Regen. 2021;29(4):642-9. https://doi.org/10.1111/wrr.12939
43. Huang C-H, Yang T-T, Lin K-I. Mechanisms and functions of SUMOylation in health and disease: a review focusing on immune cells. J Biomed Sci. 2024;31(1):16. https://doi.org/10.1186/s12929-024-01003-y
44. Singha SP, Memon S, Bano U, Isaac AD, Shahani MY. Evaluation of p21 expression and related autism-like behavior in Bisphenol-A exposed offspring of Wistar albino rats. Birth Defects Res. 2022;114(11):536–50. https://doi.org/10.1002/bdr2.2022
45. Singha SP, Memon S, Kazi SAF, Nizamani GS. Gamma aminobutyric acid signaling disturbances and altered astrocytic morphology associated with Bisphenol A induced cognitive impairments in rat offspring. Birth Defects Res. 2021;113(12):911–24. https://doi.org/10.1002/bdr2.1886
Downloads
Published
Issue
Section
License
Copyright (c) 2026 Kashaf Asghar, Rameesha Ashraf, Muhammad Taimoor, Aqeel Ahmed, Hamdan Khalid (Author)

This work is licensed under a Creative Commons Attribution-NonCommercial 4.0 International License.





